Media Summary: In this video (recorded live in class) I give a brief introduction to next generation sequencing. I describe the technology and some ... I describe base calling for next generation sequencing This lecture is part of the course "Introduction to
Statistics For Genomics Batch Effects - Detailed Analysis & Overview
In this video (recorded live in class) I give a brief introduction to next generation sequencing. I describe the technology and some ... I describe base calling for next generation sequencing This lecture is part of the course "Introduction to The models behind RMA and fRMA explained. At the end quality assessment comes up and I show some examples of the utility of ... This lab shows some simple R commands for creating dendrograms and MDS plots. I briefly explain the biology of DNA methylation, CpG Islands and measurement technology (microarrays and sequencing).
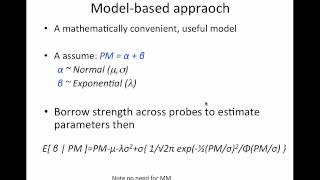
Presented at BOSC 2022, part of ISMB, in Madison, WI. In this lecture I introduce the basics of I motivate and describe the normal+exponential model used by RMA to background correct microarray intensities. I also describe ... For the first 8 minutes, we cover useful techniques like the MA plot and the volcano plot. We then go through Karl Broman's slides ... On UMAP, your cells cluster by batch — not biology.