Media Summary: Do you check your stats after each run? Do you check your spending on your banking portal? Did you ever decide on renewing a ... Curtis Huttenhower is an Associate Professor of Computational Biology and Bioinformatics at the Harvard T.H. Chan School of ... So like Adam said I'm going to be switching more over to the
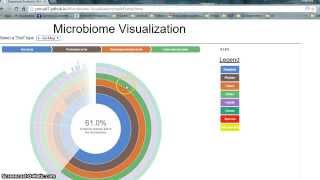
Plotcon 2017 Matthew Dillon Interactive Web Based Visualization For Microbiome Science - Detailed Analysis & Overview
Do you check your stats after each run? Do you check your spending on your banking portal? Did you ever decide on renewing a ... Curtis Huttenhower is an Associate Professor of Computational Biology and Bioinformatics at the Harvard T.H. Chan School of ... So like Adam said I'm going to be switching more over to the We present the initial alpha release of QIIME 2, a Python 3 framework supporting Cultivating Communities: Making Sense of Host- Ning Mao has shown that a safe bacteria found in dairy products has the serendipitous benefit of inhibiting the progression of a ...
Welcome to the C-DEBI/EBICS/BEACON Introduction to Bioinformatics for Metagenomics In this animation, see an example of how genetically engineered microbes being developed by researchers at the Wyss Institute ... Webinar recording will familiarize you with the issues critical to producing high quality assemblies and review current state of the ... Dr. Jack Gilbert introduces a new intensive 3-week course offered at the Marine Biological Laboratory in September. Katherine Pollard, PhD, visited Stanford Medical School to deliver the Computational Immunology Seminar. She discusses how ... On February 29, 2016, Dr. Callahan delivered this talk at the annual CEHG symposium on Stanford campus. CEHG is Stanford's ...